The goal here is to run a sequence alignment program called Novoalign on each of a set of fastq files against the hg19 human genome. The alignment for each input sequence is independent of the others. So this project is well suited for being converted into a swarm of jobs where all run simultaneously, thus vastly reducing the time required to process the data.
In the previous Data Storage hands-on section, you should have copied the class scripts to your /data area. If you did not, type
hpc-classes biowulfnow. This command will copy the scripts and input files used in this online class to your /data area, and will take about 5 minutes.
Let's look at the data:
[biowulf]$ cd /data/$USER/hpc-classes/biowulf/serial2swarm [biowulf]$ ls fastq_files.tar.gz serial.sh single.sh make_swarm.sh make_swarm2.shfastq_files.tar.gz = a tar file containing a set of fastq-format files called xaa, xab etc
There are several factors to consider before running a swarm:
Let's determine the unknown factors in the list above. We're going to run a single test job and determine these values. Let's untar the fastq files -- we can determine how many there are (which will in turn determine the number of subjobs) and use one for our test.
The script 'single.sh' is the same as 'serial.sh' except it runs just one of the query sequences. If you 'diff' the two files, you'll see the changes. Submit it to the batch system, requesting 20 GB of memory, 16 CPUs, and 10 hours of walltime.
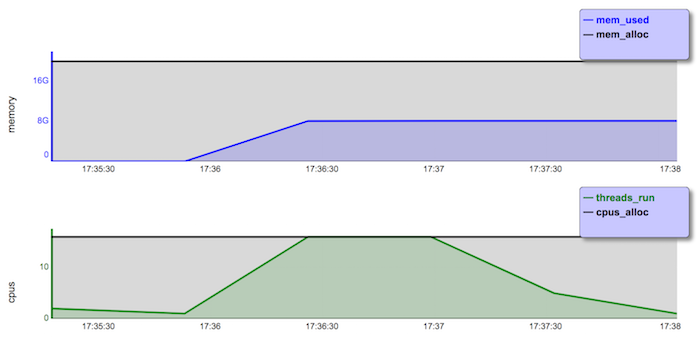
biowulf% sbatch --mem=20g --cpus-per-task=16 --time=10:00:00 single.shMonitor the job with 'squeue -u $USER', or 'jobload -u $USER' until it completes. Look at the Slurm output file and use 'jobhist jobnumber' or the user dashboard to get the information needed.
biowulf% jobhist 2555723 JobId : 2555723 User : user Submitted : 20180603 17:34:27 Started : 20180603 17:34:38 Ended : 20180603 17:37:41 Submission Path : /data/user/hpc-classes/biowulf/serial2swarm Submission Command : sbatch --mem=20g --cpus-per-task=16 --time=10:00:00 single.sh Jobid Partition State Nodes CPUs Walltime Runtime MemReq MemUsed Nodelist 2555723 norm COMPLETED 1 16 00:30:00 00:03:03 20.0GB/node 8.1GB cn3168

biowulf% ls -1 input | wc -l 90This reports that there are 90 query files in the 'input' directory. Thus, the swarm would have 90 subjobs. No problem, batchlim reports that the max array size (max number of subjobs in a swarm) is 1001.
Input files: 6.6 GB
Space for a single output file: 70MB
Space for all 90 output files: 70*90 = 6300 MB = 6.3 GB
So the total space needed for this project would be 6.6 + 6.3 GB = 13 GB. Use checkquota or the User Dashboard to make sure you have at least 7 GB free space in /data/$USER before submitting this swarm.
In future, if you planned to run a large swarm of, say, 1000 such alignments similar to these, you would know that you needed at least ~150 GB of disk space to store the input and output files. If your current /data area did not have enough space and you cannot clear out unused files, you should request more space, using the numbers you calculated here as a justification.
Next we need to create a swarm command file that contains one line for each Novoalign alignment run. The script make_swarm.sh is a modification of the serial.sh script that simply prints out the Novoalign command for each input query sequence into a file.
biowulf% sh make_swarm.sh working on file xaa working on file xab working on file xac [...]After running this script, you should see the file novo_swarm.sh in the directory, containing lines like this:
novoalign -c $SLURM_CPUS_PER_TASK -f input/xaa -d /fdb/novoalign/chr_all_hg19.nbx -o SAM > out/xaa.sam novoalign -c $SLURM_CPUS_PER_TASK -f input/xab -d /fdb/novoalign/chr_all_hg19.nbx -o SAM > out/xab.sam novoalign -c $SLURM_CPUS_PER_TASK -f input/xac -d /fdb/novoalign/chr_all_hg19.nbx -o SAM > out/xac.sam [...]This is the swarm command file.
Now that we have a swarm command file and the swarm parameters we need, we can submit the swarm.
biowulf% swarm -f novo_swarm.sh -g 10 -t 8 --time=10:00 --module novocraftwhere
| -f novo_swarm.sh | swarm command file |
| -g 10 | 10 GB of memory per subjob, i.e. per alignment |
| -t 8 | 8 CPUs per subjob. Novoalign will run 8 threads on the allocated 8 CPUs, because of the -c $SLURM_CPUS_PER_TASK parameter in the swarm command file. If you decide to run 4 or 16 threads instead, you only need to change the -t parameter when submitting the swarm. The swarm command file can remain unchanged. |
| --time=10:00 | walltime of 10 mins per subjob |
Use 'sjobs' or 'squeue' to see if the jobs are running.
biowulf% sjobs User JobId JobName Part St Reason Runtime Walltime Nodes CPUs Memory Dependency Nodelist ================================================================================================================ user 2556165_0 swarm norm R --- 1:08 10:00 1 8 10GB/node cn3139 user 2556165_1 swarm norm R --- 1:08 10:00 1 8 10GB/node cn3176 [..]
Use 'jobload' to check the loads and memory:
biowulf% jobload
JOBID TIME NODES CPUS THREADS LOAD MEMORY
Elapsed / Wall Alloc Active Used / Alloc
2556165_89 00:02:24 / 00:10:00 cn3348 8 8 100% 1.7 / 10.0 GB
2556165_0 00:02:25 / 00:10:00 cn3139 8 8 100% 0.4 / 10.0 GB
2556165_2 00:02:25 / 00:10:00 cn3345 8 8 100% 7.9 / 10.0 GB
2556165_3 00:02:25 / 00:10:00 cn3344 8 8 100% 7.9 / 10.0 GB
[..]
Looks good! Jobs that have just started up may not show the full memory usage yet, but eventually all the subjobs should show 100% CPU utilization and ~8 GB of memory used.
After the jobs complete, use 'jobhist' to see how long they took.
biowulf% jobhist 2556165
JobId : 2556165
User : user
Submitted : 20180603 18:31:19
Started : 20180603 18:31:40
Ended : 20180603 18:36:09
Submission Path : /data/user/hpc-classes/biowulf/serial2swarm
Submission Command : sbatch --array=0-89 --job-name=swarm --output=/data/user/hpc-
classes/biowulf/serial2swarm/swarm_%A_%a.o --error=/data/user
/hpc-classes/biowulf/serial2swarm/swarm_%A_%a.e --cpus-per-task=8
--mem=10240 --partition=norm --time=10:00
/spin1/swarm/user/2556165/swarm.batch
Swarm Path : /data/user/hpc-classes/biowulf/serial2swarm
Swarm Command : /usr/local/bin/swarm -f novo_swarm.sh -g 10 -t 8 --time=10:00
--module novocraft
In this case, all the alignments finished within 5 mins. If they had been run sequentially as in the serial.sh script, the alignments would have taken 90*3 = 270 mins =~ 4.5 hours.
This is left as an exercise. Examine the file make_swarm2.sh. It performs the same tasks as we did manually above
Try it!
biowulf% sh make_swarm2.sh Space taken at start 1.6G/data/teacher/hpc-classes/biowulf/serial2swarm Unpacking input fastq files Space taken for input files 6.6G/data/teacher/hpc-classes/biowulf/serial2swarm 2558027