AMPL is the Accelerating Therapeutics for Opportunites in Medicine (ATOM) Consortium Modeling PipeLine for drug discovery.
Allocate an interactive session and run the program.
Sample session (user input in bold):
[user@biowulf ~]$ sinteractive --gres=lscratch:10 --mem=8g -c4 --tunnel
salloc: Pending job allocation 35296239
salloc: job 35296239 queued and waiting for resources
salloc: job 35296239 has been allocated resources
salloc: Granted job allocation 35296239
salloc: Waiting for resource configuration
salloc: Nodes cn0857 are ready for job
srun: error: x11: no local DISPLAY defined, skipping
error: unable to open file /tmp/slurm-spank-x11.35296239.0
slurmstepd: error: x11: unable to read DISPLAY value
Created 1 generic SSH tunnel(s) from this compute node to
biowulf for your use at port numbers defined
in the $PORTn ($PORT1, ...) environment variables.
Please create a SSH tunnel from your workstation to these ports on biowulf.
On Linux/MacOS, open a terminal and run:
ssh -L 36858:localhost:36858 user@biowulf.nih.gov
For Windows instructions, see https://hpc.nih.gov/docs/tunneling
[user@cn0857 ~]$ module load ampl
[+] Loading ampl 1.3.0 on cn0857
[user@cn0857 ~]$ ipython
Python 3.7.10 (default, Feb 26 2021, 18:47:35)
Type 'copyright', 'credits' or 'license' for more information
IPython 7.16.1 -- An enhanced Interactive Python. Type '?' for help.
In [1]: import atomsci
In [2]: quit()
[user@cn0857 ~]$ cd /lscratch/$SLURM_JOB_ID
[user@cn0857 35296239]$ cp -r ${AMPL_HOME}/ampl .
[user@cn0857 35296239]$ cd ampl/atomsci/ddm/
[user@cn0857 ddm]$ jupyter-notebook --no-browser --port=$PORT1
[I 14:23:09.610 NotebookApp] Serving notebooks from local directory: /lscratch/35296239/ampl/atomsci/ddm
[I 14:23:09.610 NotebookApp] Jupyter Notebook 6.4.10 is running at:
[I 14:23:09.610 NotebookApp] http://localhost:36858/?token=5ebfcff3cfccd637d36250f9a5a8191a22011edd2a262db5
[I 14:23:09.610 NotebookApp] or http://127.0.0.1:36858/?token=5ebfcff3cfccd637d36250f9a5a8191a22011edd2a262db5
[I 14:23:09.610 NotebookApp] Use Control-C to stop this server and shut down all kernels (twice to skip confirmation).
[C 14:23:09.618 NotebookApp]
To access the notebook, open this file in a browser:
file:///spin1/home/linux/user/.local/share/jupyter/runtime/nbserver-63375-open.html
Or copy and paste one of these URLs:
http://localhost:36858/?token=5ebfcff3cfccd637d36250f9a5a8191a22011edd2a262db5
or http://127.0.0.1:36858/?token=5ebfcff3cfccd637d36250f9a5a8191a22011edd2a262db5
Now in a new terminal on your local machine, you should initiate a new ssh session to Biowulf to establish the ssh tunnel. Using the port that was allocated to you during the sinteractive step above:
[user@mymachine ~]$ ssh -L 36858:localhost:36858 user@biowulf.nih.gov [user@biowulf ~]$
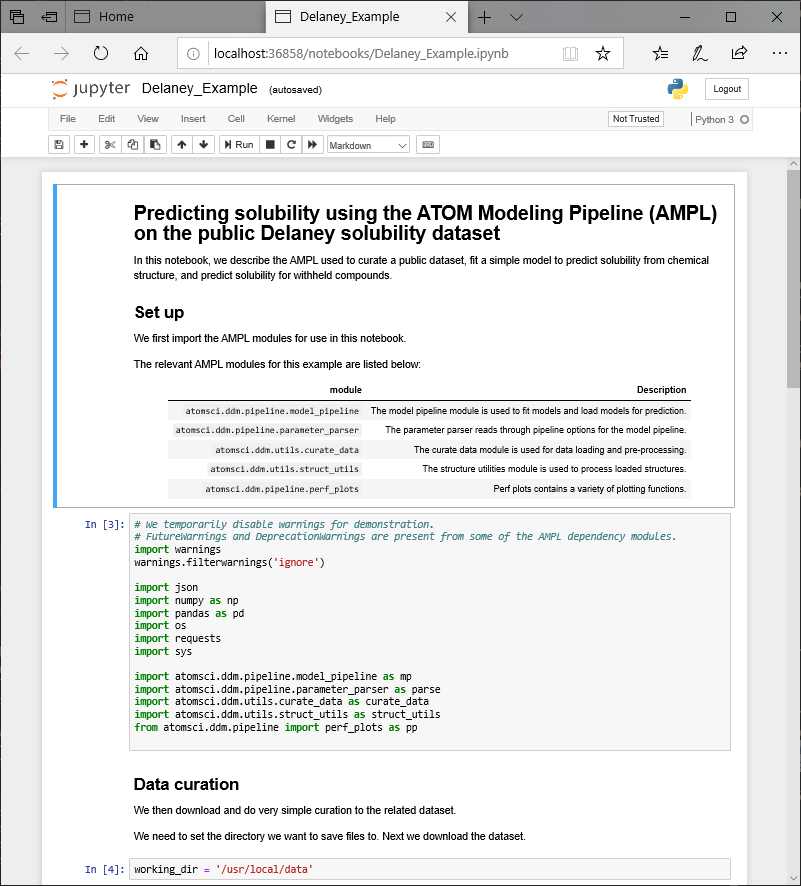
This will allow you to open a local browser and view the Jupyter Notebook session running on the compute node:

^C[I 11:48:01.862 NotebookApp] interrupted Serving notebooks from local directory: /lscratch/1151805/ampl/atomsci/ddm 1 active kernel The Jupyter Notebook is running at: http://localhost:36858/?token=05982663a539c9fac05608e4b7ca76649356f1cdb3dcc750 Shutdown this notebook server (y/[n])? y [C 18:43:10.271 NotebookApp] Shutdown confirmed [I 11:48:03.637 NotebookApp] Shutting down 1 kernel [I 11:48:03.838 NotebookApp] Kernel shutdown: 1b0c8fe9-a2de-45bc-ab5a-bea16e60dc0f [user@cn0859 ddm]$ exit exit salloc.exe: Relinquishing job allocation 1151805
Create a batch input file (e.g. ampl.sh). For example:
#!/bin/bash set -e module load ampl # command here
Submit this job using the Slurm sbatch command.
sbatch [--cpus-per-task=#] [--mem=#] ampl.sh
Create a swarmfile (e.g. ampl.swarm). For example:
cmd1 cmd2 cmd3 cmd4
Submit this job using the swarm command.
swarm -f ampl.swarm [-g #] [-t #] --module amplwhere
| -g # | Number of Gigabytes of memory required for each process (1 line in the swarm command file) |
| -t # | Number of threads/CPUs required for each process (1 line in the swarm command file). |
| --module ampl | Loads the ampl module for each subjob in the swarm |