setwd("/data/somewhere")
Another way (more recommended) is using project, which will set it's own working directory. File -> New Project -> New Directory -> Create New Project (assign directory name, and choose the directory location such as "/data/XXX/")i
QStandardPaths: XDG_RUNTIME_DIR not set, defaulting to '/XXXX/'
libGL error: No matching fbConfigs or visuals found
libGL error: failed to load driver: swrast
Unrecognized OpenGL version
or
libGL error: failed to load driver: swrast
Unrecognized OpenGL version
qt.qpa.xcb: could not connect to display
qt.qpa.plugin: Could not load the Qt platform plugin "xcb" in "" even though it was found.
qt.qpa.plugin: Could not load the Qt platform plugin "xcb" in "" even though it was found.
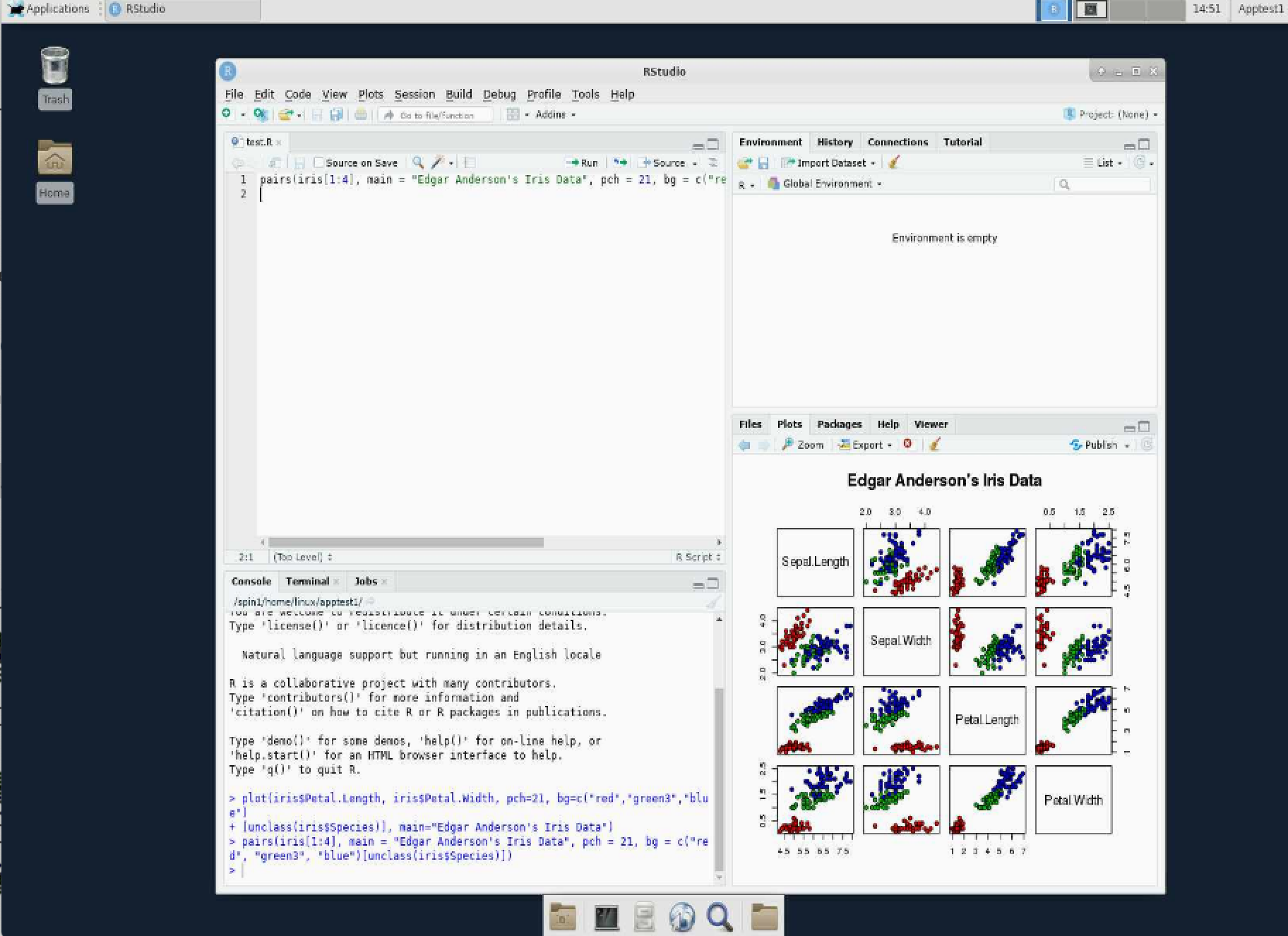
Running RStudio interactively
Connect to Biowulf using a Graphical Session on HPC OnDemand, then open a terminal:


Allocate an sinteractive session. It is generally recommended to allocate at least a small amount of lscratch for temporary storage for R.
[user@biowulf ~]$ sinteractive --mem=10g --gres=lscratch:5 salloc.exe: Pending job allocation 15323416 salloc.exe: job 15323416 queued and waiting for resources salloc.exe: job 15323416 has been allocated resources salloc.exe: Granted job allocation 15323416 salloc.exe: Waiting for resource configuration salloc.exe: Nodes cn1640 are ready for job [user@cn1640 ~]$ module load rstudio R [+] Loading Rstudio 1.1.447 Remember to load an R module before starting Rstudio ... [user@cn1640 dir]$ rstudio &

Documentation
